Hello,
I found that the Xlink.pl script wrote by Jason DeJoannis, Stephen Todd and James Wescott, etc is very useful in building the crosslinked structure of thermosets, especially the epoxy resin. Recently I tried to build a crosslinked network of phenolic resin, which mainly consist of phenols (PhOH) and methlyenes (-CH2-) (see Fig. 1).
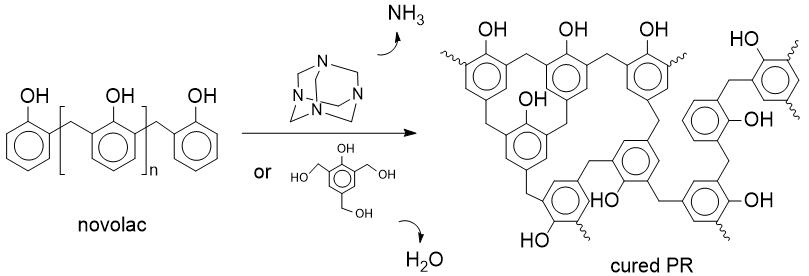
Fig. 1 The cure process of phenolic resin.
I built the following initial models and crosslinked them with Xlink.pl,
Oligomer: octamer (PhOH-(PhOHCH2)6-PhOH), the 2, 4 and 6 sites are named as R1
Linker: Tri-methyl PhOH, the C atom named as R2 and not a multi-linker
Result → End with an Error “There are no objects in the "Molecules" collection (function/property "Molecule") at -e line 1006”.
I checked the line 1006 and can't find any mistake,
I would like to know whether these strategies are right.
Any suggestions about the model construction method or the way to modify the script will be helpful.
Best,
Tony Cheng
I found that the Xlink.pl script wrote by Jason DeJoannis, Stephen Todd and James Wescott, etc is very useful in building the crosslinked structure of thermosets, especially the epoxy resin. Recently I tried to build a crosslinked network of phenolic resin, which mainly consist of phenols (PhOH) and methlyenes (-CH2-) (see Fig. 1).
Fig. 1 The cure process of phenolic resin.
I built the following initial models and crosslinked them with Xlink.pl,
Oligomer: octamer (PhOH-(PhOHCH2)6-PhOH), the 2, 4 and 6 sites are named as R1
Linker: Tri-methyl PhOH, the C atom named as R2 and not a multi-linker
Result → End with an Error “There are no objects in the "Molecules" collection (function/property "Molecule") at -e line 1006”.
I checked the line 1006 and can't find any mistake,
- # Rename the xlinker and oligomer molecules at the beginning
- if (\\\$row == 0)
- {
- foreach my \\\$atom (@{\\\$doc1->UnitCell->Atoms})
- {
- if (\\\$atom->Name eq "\\\$xlinkerReactiveAtom")
- {
- \\\$atom->Ancestors->Molecule->Name = \\\$xlinkerName;
- }
- elsif (\\\$atom->Name eq "\\\$oligomerReactiveAtom")
- {
- \\\$atom->Ancestors->Molecule->Name = \\\$oligomerName;
- }
- }
I would like to know whether these strategies are right.
Any suggestions about the model construction method or the way to modify the script will be helpful.
Best,
Tony Cheng

