I am modeling the nature rubber and I use the Materials Studio 5.0 to built the rubber system. However, when I want to built the confined model of the rubber, the job is always failed (see the attachment). Is there anything wrong with me method?
1) I use the c_isoprene in the lib to generate a polymer chain in Homopolymer part.
2) I use the generated chain to build the polymer structure in amorphous cell.
Below is the .out file for this job.
DISCOVER Molecular Simulation Program
Version: 2009.1
Build: Oct 22 2009
Date: Thu Mar 11 20:20:07 2010
Host Name: TAM04
Host Type: Windows
Threads: 1
---------------------------------------------------------------
Checked out license feature: MS_discover_MP [for LSD] (1 copy)
---------------------------------------------------------------
randomSeed is set to 360418
Line 5:BTCL> autoEcho off
---------------------------------------------------------------
Checked out license feature: MS_compass_MP [for LSD] (1 copy)
---------------------------------------------------------------
INPUT FILES
___________
File Type Name
--------- ----
Forcefield C:\\PROGRA~1\\Accelrys\\MATERI~1.0\\etc\\Gateway\\..\\..\\share\\Discover\\res/compass.frc
Molecular data Polyc_isoprene.mdf
Coordinate Polyc_isoprene.car
Periodic Boundary Conditions
____________________________
Length (A) Angle (degrees)
---------- ---------------
a 36.91000 alpha 90.00000
b 36.91000 beta 90.00000
c 36.91700 gamma 90.00000
******************************
Processing frame number 1
******************************
MOLECULAR TOPOLOGY
__________________
Number of molecules: 10
Number of residues: 400
Number of atoms: 5220 (asymmetric unit: 5220)
Number of atom types: 4
Number of bonds: 5210
Number of consolidated angles: 9600
Number of consolidated torsions: 12310
Number of bond_bond_1_3s: 12310
Number of angle-angles: 16800
Number of out-of-planes: 800
FORCEFIELD OPTIONS
__________________
Filename : compass.frc
Definition name : cff91
Version : 2.8
Last modification date : 3/6/2007
# of automatic parameters : 0
NONBOND ENERGY CUTOFFS
______________________
Cutoff (A) Spline Width (A) Buffer Width (A)
vdW 8.50 0.00 0.50
Coulomb -- -- --
NONBOND ENERGY CMM PARAMETERS
_____________________________
Number of Cells Update Width (A)
Coulomb 1 1.00
Summation method for vdW : Atom based
Summation method for Coulomb : Cell multipole
Dielectric : 1.00
Dynamics: Warning: Stress (pressure) calculation not performed since the nonbond method used does not calculate the Virial (cell derivatives)
MOLECULAR DYNAMICS
__________________
Ensemble : NVT
Temperature : 298.00 K
Control Method : Direct Velocity Scaling, Temp. Window = 10.00 K
Timestep : 1.00 fs
Duration : 2000.00 fs
Integration Method : Velocity Verlet
Initial Velocities : Random Velocities from Boltzmann distribution
Initial Temp. : 298.00 K
Error: Dynamics: Energy difference between successive steps in dynamics is greater than the user defined variable DEVIATION, 5000.000
invoked from within
"dynamics time = 2000 timestep = 1.0 initial_temperature = 298.00 +boltzmann ensemble = NVT temperature_control_method = velocity_scaling integrati ..."
("while" body line 10)
invoked from within
"while {[readFile archive frame = \\\$iframe filename = \\\$arcname] != ""} {
echo "\\n******************************"
echo "Processing fr ..."
Total time used by DISCOVER: 1.47 secs
Completion date: Thu Mar 11 20:20:25 2010
Exiting Discover: An ERROR has occurred!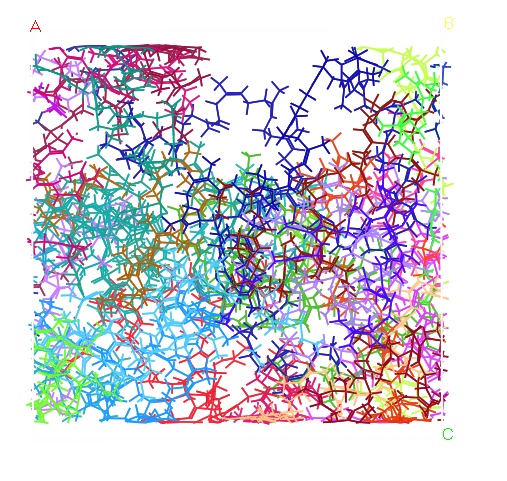
1) I use the c_isoprene in the lib to generate a polymer chain in Homopolymer part.
2) I use the generated chain to build the polymer structure in amorphous cell.
Below is the .out file for this job.
DISCOVER Molecular Simulation Program
Version: 2009.1
Build: Oct 22 2009
Date: Thu Mar 11 20:20:07 2010
Host Name: TAM04
Host Type: Windows
Threads: 1
---------------------------------------------------------------
Checked out license feature: MS_discover_MP
---------------------------------------------------------------
randomSeed is set to 360418
Line 5:BTCL> autoEcho off
---------------------------------------------------------------
Checked out license feature: MS_compass_MP
---------------------------------------------------------------
INPUT FILES
___________
File Type Name
--------- ----
Forcefield C:\\PROGRA~1\\Accelrys\\MATERI~1.0\\etc\\Gateway\\..\\..\\share\\Discover\\res/compass.frc
Molecular data Polyc_isoprene.mdf
Coordinate Polyc_isoprene.car
Periodic Boundary Conditions
____________________________
Length (A) Angle (degrees)
---------- ---------------
a 36.91000 alpha 90.00000
b 36.91000 beta 90.00000
c 36.91700 gamma 90.00000
******************************
Processing frame number 1
******************************
MOLECULAR TOPOLOGY
__________________
Number of molecules: 10
Number of residues: 400
Number of atoms: 5220 (asymmetric unit: 5220)
Number of atom types: 4
Number of bonds: 5210
Number of consolidated angles: 9600
Number of consolidated torsions: 12310
Number of bond_bond_1_3s: 12310
Number of angle-angles: 16800
Number of out-of-planes: 800
FORCEFIELD OPTIONS
__________________
Filename : compass.frc
Definition name : cff91
Version : 2.8
Last modification date : 3/6/2007
# of automatic parameters : 0
NONBOND ENERGY CUTOFFS
______________________
Cutoff (A) Spline Width (A) Buffer Width (A)
vdW 8.50 0.00 0.50
Coulomb -- -- --
NONBOND ENERGY CMM PARAMETERS
_____________________________
Number of Cells Update Width (A)
Coulomb 1 1.00
Summation method for vdW : Atom based
Summation method for Coulomb : Cell multipole
Dielectric : 1.00
Dynamics: Warning: Stress (pressure) calculation not performed since the nonbond method used does not calculate the Virial (cell derivatives)
MOLECULAR DYNAMICS
__________________
Ensemble : NVT
Temperature : 298.00 K
Control Method : Direct Velocity Scaling, Temp. Window = 10.00 K
Timestep : 1.00 fs
Duration : 2000.00 fs
Integration Method : Velocity Verlet
Initial Velocities : Random Velocities from Boltzmann distribution
Initial Temp. : 298.00 K
Error: Dynamics: Energy difference between successive steps in dynamics is greater than the user defined variable DEVIATION, 5000.000
invoked from within
"dynamics time = 2000 timestep = 1.0 initial_temperature = 298.00 +boltzmann ensemble = NVT temperature_control_method = velocity_scaling integrati ..."
("while" body line 10)
invoked from within
"while {[readFile archive frame = \\\$iframe filename = \\\$arcname] != ""} {
echo "\\n******************************"
echo "Processing fr ..."
Total time used by DISCOVER: 1.47 secs
Completion date: Thu Mar 11 20:20:25 2010
Exiting Discover: An ERROR has occurred!

