Hi,
I just want to make calcium glycolate (its a salt of glycolic acid.) and optimized it using Pcff.
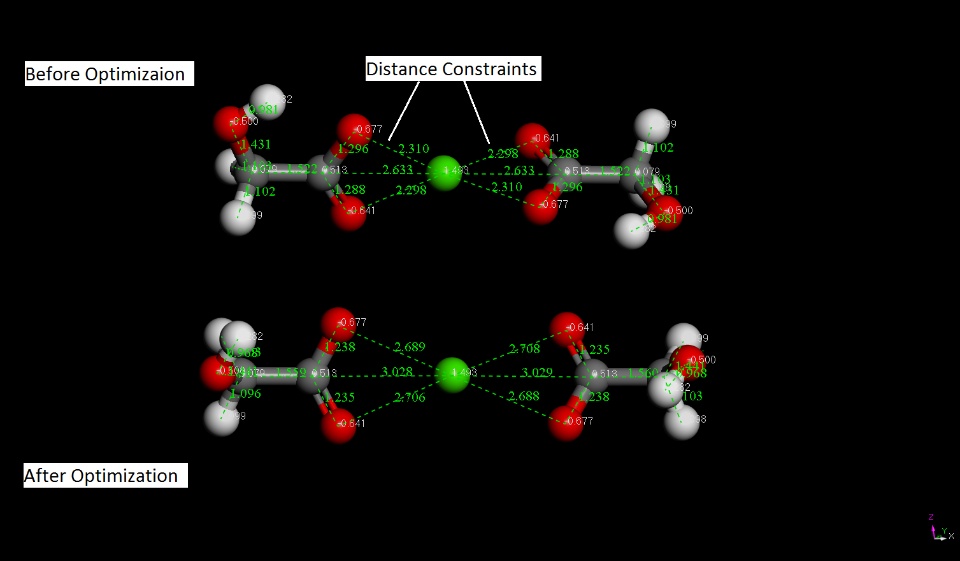
I am trrying to create calcium glycolate moleucle in the MS7. I assined dft charges to the molecule and then contrained distance between O- (anion) and Ca++ (cation) [refer the image]. Also, I selected auto force field assigneent and use current charge option.
I am able to see that ditance is fixed before geometry optimization. (There was tick mark in the distance contraint column.) Howvever, after pcff optimzation, distance between calcium ion and oxygen anion got chnage.
How can I constrant the distances? is there another way to make a calcium glycolate molecule ?
Thank you for the help.
I just want to make calcium glycolate (its a salt of glycolic acid.) and optimized it using Pcff.
I am trrying to create calcium glycolate moleucle in the MS7. I assined dft charges to the molecule and then contrained distance between O- (anion) and Ca++ (cation) [refer the image]. Also, I selected auto force field assigneent and use current charge option.
I am able to see that ditance is fixed before geometry optimization. (There was tick mark in the distance contraint column.) Howvever, after pcff optimzation, distance between calcium ion and oxygen anion got chnage.
How can I constrant the distances? is there another way to make a calcium glycolate molecule ?
Thank you for the help.

