Hello,
I'am using Babel and Corina (from molecular-networks.com) to convert form a LPS structure form a chemdraw file to a pdb format (.cdw -> (babel) -> simles/sdf -> (corina) -> pdb.
What i get in the end is a molecule with a single chain and a single residue (UNK1) with all the atoms. How can i select groups of atoms and define them as a new residue (ex: GLC, GAL, FUC, ..etc ) within the molecule. I want to get something like 1CAP where each individual monosaccharide is defined in an idependent residue.
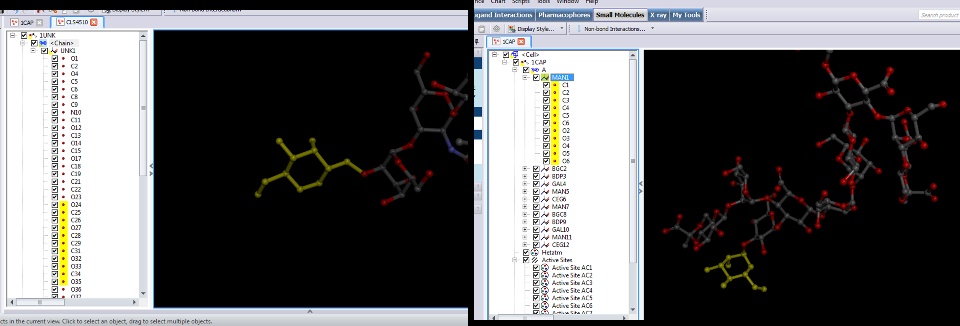
I'am using Babel and Corina (from molecular-networks.com) to convert form a LPS structure form a chemdraw file to a pdb format (.cdw -> (babel) -> simles/sdf -> (corina) -> pdb.
What i get in the end is a molecule with a single chain and a single residue (UNK1) with all the atoms. How can i select groups of atoms and define them as a new residue (ex: GLC, GAL, FUC, ..etc ) within the molecule. I want to get something like 1CAP where each individual monosaccharide is defined in an idependent residue.

